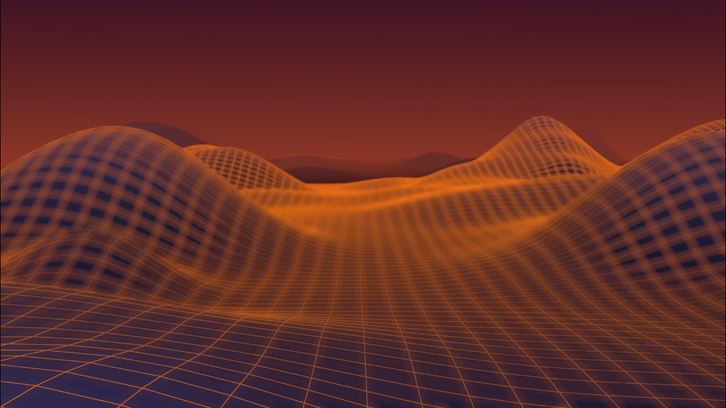
3D Animation
Protein Fitness Landscapes
and Directed Evolution
Story
Protein fitness landscapes and directed evolution are fundamental concepts in the field of protein engineering. Although the concepts themselves are highly dimensional, they are often introduced to students using multiple 2D representations rendered from data visualization software. This animation was created to help instructors explain these concepts, and their underlying components, in a continuous and cohesive story that captures attention and inspires exploration.
Details
Client: Phillip Romero, BSE, PhD,
Department of Biochemistry,
University of Wisconsin-Madison
Audience: Undergraduate/Graduate level students
Software: 3ds Max, rendered with VRay, Visual Molecular Dynamics (VMD, ADobe Photoshop, Illustrator, After Effects, and Audition
Final Animation
Research
This project started with a zoom conversation between Dr. Romero and I, where he walked me through the typical lecture that he gives his students on protein evolution and the concepts of Protein Sequence Space, Protein Fitness Landscapes, and Directed Evolution. After some discussion I was able to pinpoint hurdles to his students' understanding of these concepts and their relationships and we both agreed that they were a suitable subject for a 3D animation. After our meeting I started combing through literature to familiarize myself with each concept and the visualizations currently in use. Below is an organizational chart with some of my background research:
Script & Storyboard
Once I felt comfortable with the material, I began narrowing the scope of information and thinking about how the linear story that Dr. Romero tells using current 2D representations would flow in 3D space. After an initial script and storyboard review over zoom, Dr. Romero provided me with feedback that I incorporated into the final storyboard and narration below:
Models
The protein that I used to represent non-specific proteins in this animation was a somewhat simple-shaped protein that Dr. Romero's lab is working with and was well defined in the Protein Data Bank (PDB ID: 2QMT). I pulled the Quick Surf and ribbon models from VMD to serve as the main protein structure throughout the animation. The other major model in the animation, the fitness landscape, I created by soft selected vertices and applying a UVW map for the heatmap effect.
Look and Feel
Throughout the literature, and within Dr. Romero's lecture, I found that protein fitness landscapes and directed evolution are described using phrases like "walking", "traversing", "traveling", or "stepping" and the graphical representations of these landscapes are physically described as having varying severity of "terrains". Since there is already so much emphasis on the landscape metaphor, I wanted to evoke the feeling of an expansive terrain and have the viewer feel like they're physically moving through this space. I drew a lot of inspiration from landscape photography and aerial fly-overs to direct my camera movement.
For the color pallet and style of the animation I decided that using highly saturated points of light would be an effective method to direct the viewer's attention and cue certain actions in the scene. The evolutionary steps moving up to the fitness peak, for example, were inspired by bioluminescent algae being disrupted in water. This style also fits well with the concept of moving from a natural process to data driven representation.


Animation
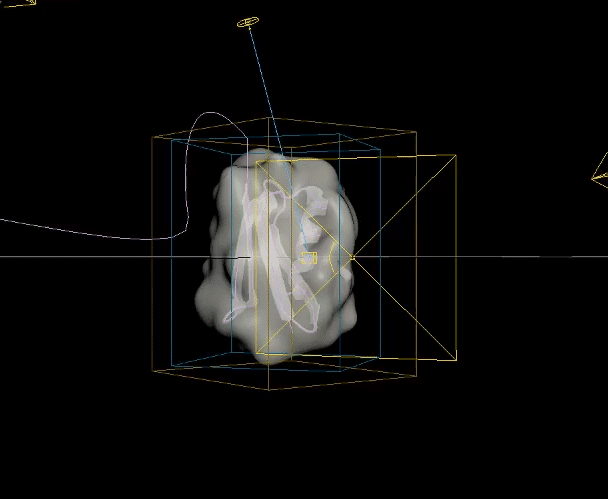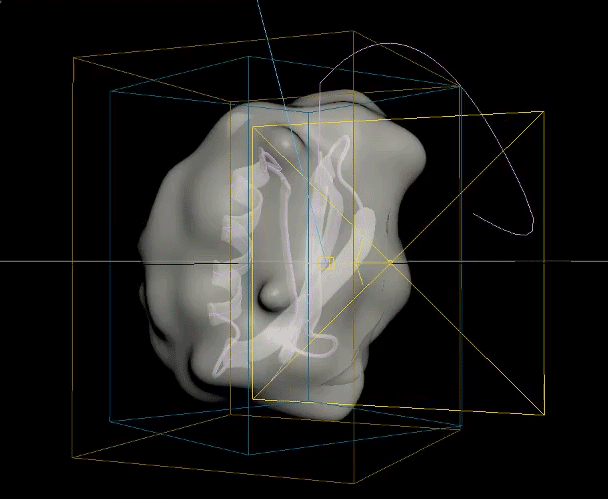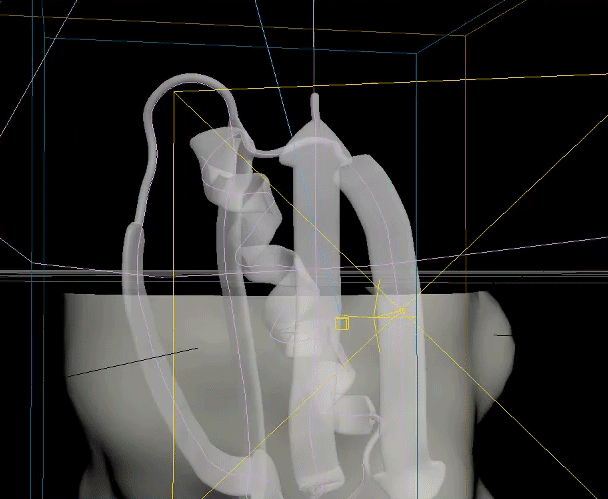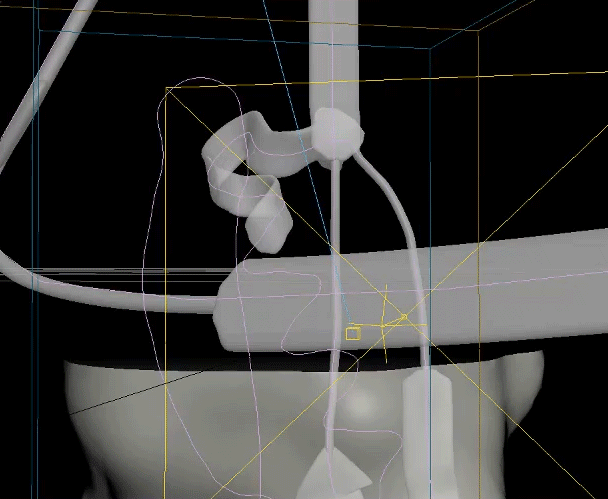

The two biggest challenges in this animation were creating the unfolding ribbon model and amino acid odometer effects. To create the unfolding effect I used a spline created from the original ribbon model, plus a tail leading off the frame, and added a follow path constraint to a simple box model of the ribbon so it folded along the spline and could be animated moving along it. For the odometer effect, I was unable to find a suitable plugin or code to produce the effect with letters rather than numbers, so I hand animated columns of text to simulate the ticking of a mechanical counter.
In addition to those two effects, I incorporated several 2D and pseudo-2D elements including the fitness line for the graph and path along the peak. These were created by selecting edges on the landscape model and converting the selection to splines that could then be rendered out in scanline and composited back in.
Composited Stills
The final result was meant to emulate familiar 2D visualizations currently used by the protein engineering community, while providing a cohesive, linear, and visually compelling story.
Feedback
After the completion of this project I met with Dr. Romero and he provided final feedback and comments that will be incorporated into a later version of the animation